Xinzeng Feng
The mechanical forces arising from the interaction between a cell and the extracellular matrix (ECM) are known to regulate cellular functions, including differentiation, growth, adhesion, and migration. Traditionally, cell-generated forces are measured when cells are plated on 2D substrate. However, in vivo, cells are mostly embedded within a 3D construct. This brings the need to quantify cellular traction forces in 3D. In vitro experiments, a dense population of fluorescent microbeads was embedded within the matrix. Upon release of cell forces, gel deformation was measured by the bead displacement, which I used to infer cellular traction forces in conjunction with the known gel properties. In specific, using a technique called Cell Traction Microscopy, I determined the 3D traction distribution by solving an inverse problem in the ECM which is assumed to be linearly elastic. With the development of more accurate mechanic models of ECM, this technique provides a promising tool to study cell-ECM interaction in a quantitative way. Moving forward, my first project at The University of Texas at Austin is focused on incorporating tissue mechanical properties into tumor modeling. The goal is to develop a better mechanically coupled tumor model to predict in vivo tumor growth via quantitative magnetic resonance imaging data.
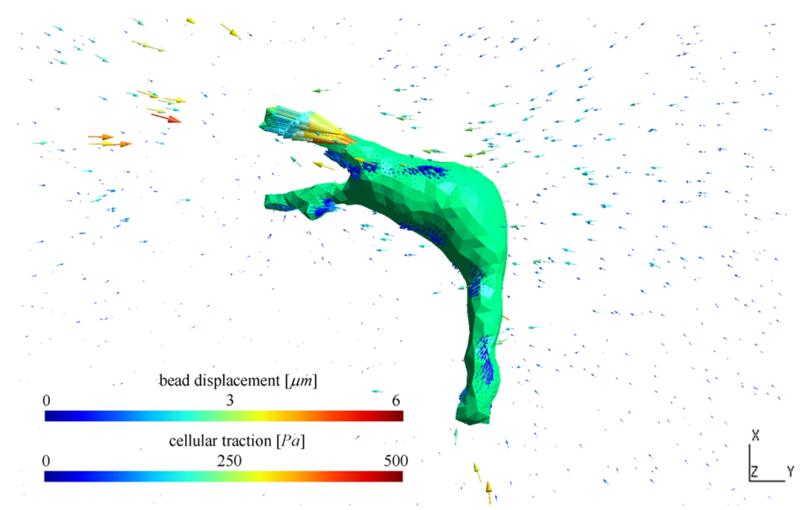
Figure. The distribution of traction forces of an MDA-MB-231 cancer cell (middle) cultured in a 2.7mg/mL unglycated type-I collagen matrix (with Young’s modulus equal to 100Pa, Poisson’s ratio equal to 0.2). The arrows on the cell surface represent the cellular traction computed using inverse method cell traction microscopy and the arrows around the cell are the measured displacement of fluorescent beads. In the calculation, force and moment balance of the cellular traction is enforced.