David Fuentes
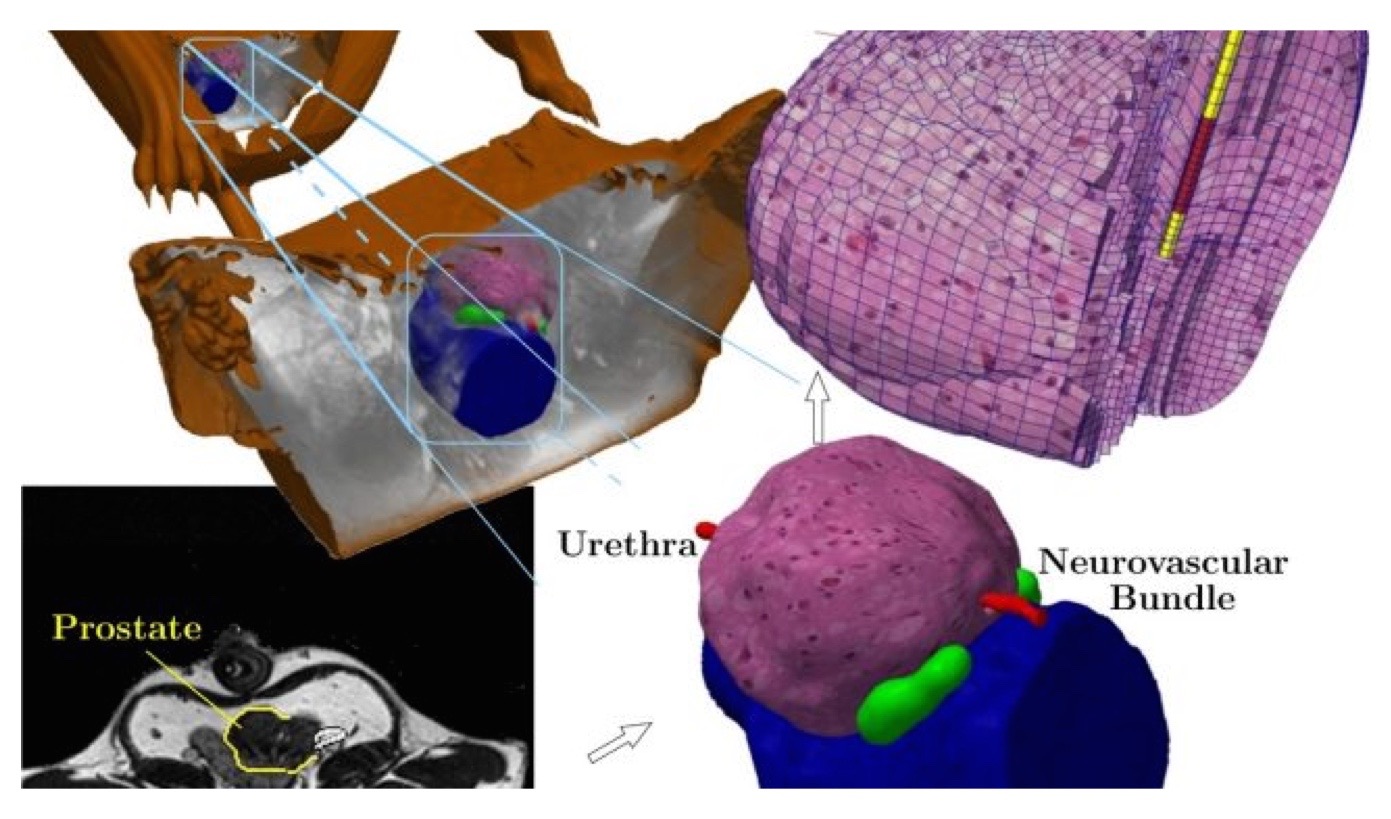
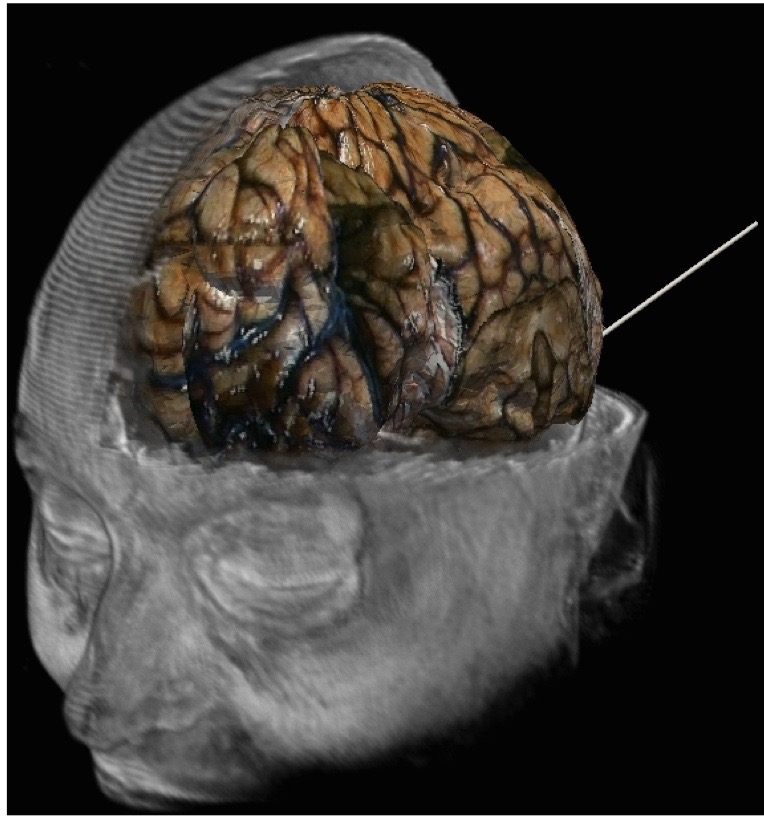
My research motivations are centered on the development of computational tools capable `predictive simulation' with applications to medical image guided diagnosis and therapy. Here, predictive simulation refers to the development of computing devices that enable confident treatment decisions from the outcome of the computer predictions. These are very ambitious research goals that embody a methodological and mathematically precise framework of computer model selection, calibration, and validation. Within this paradigm, computer models become an integral part of the therapy and image reconstruction that enable approaches previously not considered safe or possible. Given imprecisions and errors generally associated with computer predictions, predictive simulations must be performed within a probabilistic mathematical setting commonly seen within the statistics, data assimilation, and uncertainty quantification (UQ) communities. The ideas of uncertainty quantification and data assimilation are applicable to predictive computer modeling and quantitative imaging for a broad range of image guided therapies and imaging modalities, including stereotactic radiosurgery, hyperthermia, focused ultrasound, nanotechnology mediated drug therapy, and quantitative PET standard uptake values.
My research team embraces the open-science ideology for repeatibility and reproducibility. Research codes are provided on open source hosting sites, such as github (https://github.com/ImageGuidedTherapyLab), to facilitate collaborations within the lab as well as researchers within the institution. Self contained usage examples with representative data is provided as a starting point to facility reproducibility of the results. Our coding projects leverage several state of the art and actively developed software packages such as Slicer, DAKOTA, PETSc, ITK, and libMesh. This approach will facilitate collaborations with top level researchers in scientific computing on a state and national scale.